Recently I came across a nice video showing the difference between both database.
It’s a clear win for PostgreSQL.
Category: Code
MinGW / MSYS tip of the day
If ever you encounter error like this:
(159/159) checking for file conflicts [###################################################################] 100%
error: failed to commit transaction (conflicting files)
mingw-w64-i686-glib2: /mingw32/bin/gi-compile-repository.exe exists in filesystem
mingw-w64-i686-glib2: /mingw32/bin/gi-decompile-typelib.exe exists in filesystem
mingw-w64-i686-glib2: /mingw32/bin/gi-inspect-typelib.exe exists in filesystem
mingw-w64-i686-glib2: /mingw32/bin/libgirepository-2.0-0.dll exists in filesystem
...and so on...The use the following command to update your system:
pacman -Suuy --overwrite "*" KAFKA: raw write your own encoded AVRO event
Like they said, if you use a language for which serializers aren’t written yet, you can write the encoded event yourself. Here are both the recipe and the link:
| 0 | Magic Byte | Confluent serialization format version number; currently always 0. |
| 1-4 | Schema ID | 4-byte schema ID as returned by Schema Registry. |
| 5-… | Data |
https://docs.confluent.io/platform/current/schema-registry/fundamentals/serdes-develop/index.html#wire-format
#kafka #avro #schema #id #raw #event #wire #format
Kafka symbol lookup error, undefined symbol: rd_kafka_message_leader_epoch
Sometimes when using a program linked with librdkafka, you can encounter the ‘symbol lookup error.
Most of the time it’s because your program have been linked to an old version of librdkafka in your system (like outdated RedHat librdkafka yum package VS the latest build from librdkafka’s git)
Solutions:
-Check your LD_LIBRARY_PATH, be sure to find the directories containing latest librdkafka .so file is listed first
-Check what’s installed on your system
-In some case you will have to rebuild your project, having a proper LD_LIBRARY_PATH configured before
#KAFKA #C #librdkafka
GNU make, get relative PATH for i.e recursive builds with dependencies based on local git copies
In case you’re building yourself everything from source in your project, there is a good chance you once had a problem because you were using relative (like ../myressource) paths in your CFLAGS or LD_FLAGS definition and passed them down to the build system.
Short answer: when getting down in sub directories to i.e build your dependencies, the relative path can’t stay good. Only the absolute path can work.
One solution, if using GNU make, is to use something like this to compute the absolute path:
RES_RELATIVE_PATH=./path_to_my_ressource
RES_ABSOLUTE_PATH=$(realpath $(RES_RELATIVE_PATH))
CFLAGS+= -I$(RES_ABSOLUTE_PATH)
# Note: the naming is up to you. In my case, for something like cJSON, I'm using the following naming:
CJSON_RELPATH=./external/cJSON
CJSON_PATH=$(realpath $(CJSON_RELPATH))
CFLAGS+= -I$(CJSON_PATH)You can also directly initialize with a shell call:
CJSON_CFLAGS=-I$(shell realpath "./external/cJSON")
CFLAGS+= $(CJSON_FLAGS)#code #makefile #gnu #path #absolute #relative
Error 407 when using git and a proxy with authentification
Recently I stumbled upon that kind of error when trying to update a submodule:
git submodule update –remote –merge
fatal: unable to access ‘https://github.com/DaveGamble/cJSON.git/’: Received HTTP code 407 from proxy after CONNECT
Turns out that it was the proxy authentification not being sent correctly.
In our case we are using system defined proxy, so we do not need git’s ones
Solved by using:
git config –global http.proxyAuthMethod ‘basic’
git config –global –unset http.proxy
git config –global –unset https.proxy
Some also encountered SSL verification problems due to auto signed certificate use in their network, and had to disable SSL verification. It’ done with the following command, but we do not recommend to use it unless really needed:
git config –global http.sslVerify false
Correctly print uint64_t / size_t in C
Have you even stumbled upon a strange warning when trying to print an uint64_t or a size_t ?
Do no take that warning lightly, as your code can break anytime if you’re not fixing it.
Warning example:
warning: format '%llu' expects argument of type 'long long unsigned int', but argument 4 has type 'uint64_t' [-Wformat]
To avoid the warning and crashes, you have to use a one of the PRIxxx macro which will choose the right way for your system to print the value, without crashes.
Example:
#include <inttypes.h>
uint64_t i;
printf("%"PRIu64"\n", i);
Macros for printf family can be found here.
New portapack mayhem app, Level app !

The level app is as simple as possible and allow you to monitor available level meters in a single view.
There is a live graph of each RSSI values and power level.
Link to Documentation
Link to official Nightly build
or click to download compiled version at the time of the post (bin only)
#hackrf #portapack #mayhem #level-app
Edit: it’s the hundredth post of that blog, yay !
KrampusHack 2022 is opened !
KrampusHack 2022 is opened and starting on December 13 to December 31 !
goal. You are secret santa – create a surprising gift for a selected participant.
On Tuesday 13 December 2022, you will be assigned one of the other participants and you have to make your game as a gift for them (Once the competition starts, you can read here for whom you have to make a gift).
Everyone should post a wishlist of features that they would like to see in the game. Note this is a wishlist and not a requirement list – you should tune the game to your giftee, taking the wishlist as well as the person itself into account, and create something that is a nice and thoughtful surprise to them. To make your wish list easily findable, simply post it on your own log.
Rules: https://tins.amarillion.org/krampu22/rules
Join in and have fun with us ! https://tins.amarillion.org/join/
#tins #gamejam #hack #krampus #krampushack
Warnings from the compiler / linker
I was searching about a very specific warning that the gcc compiler started to sprout some few weeks ago:
That I extracted and translated from the original that was in French:
“/usr/bin/ld: warning: xxxxxxx.o: requires executable stack (because the .note.GNU-stack section is executable)”
And I stumbled upon that little gem page was one of the few to have a correct explanation:
https://www.redhat.com/en/blog/linkers-warnings-about-executable-stacks-and-segments
And also a few more here:
https://wiki.gentoo.org/wiki/Hardened/GNU_stack_quickstart
#gcc #compiler #linker #warning
Git remove commit from a pushed pull request
If ever you have to remove a commit from a pushed pull request, here is a simple way to do it. Bonus: it’s not adding a commit for the removal.
First be sure to be on the good branch
git checkout target-pull-request-branch
Then have a watch at the PRs commit tab and note the commits hash you want to remove (one or more)
Replace X by that number in the following command, which will make and interactive rebase for the X last commits
git rebase -i HEAD~X
You are now going to have an interactive text view, in which all the aimed commits are preceded by the word ‘pick’
Using the view like a text editor, replace the ‘pick’ word by ‘drop’ on the lines containing the targeted commit’s hashes .
Exemple
Replaced pick with drop for commits 4aa3149 and 2031a79b:
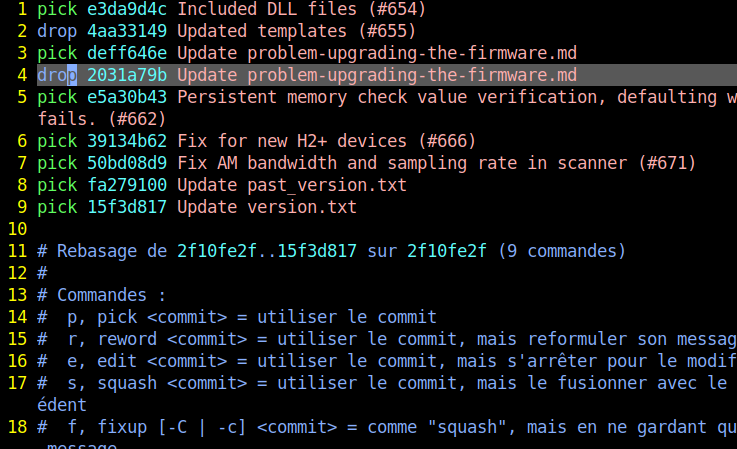
Save and exit
In my case vim was the default text editor, so here you go escape colon w q
Push force the rebased repository
git push --force
This will remove the targeted commits and the only trace will be in the PR comments. The commit page will look like the rebased one, without the discarded commits nor a mention to them.
KAFKA
KAFKA is in all mouths these days. So I share here, also as a reminder for me, a couple of #KAFKA ressources that looked helpful to me.
TLDR:
Do not use KAFKA for:
-“Little” Data flows (overkill)
-Streaming ETL (handling transformation is a hassle)
-Store and process large files (images, videos, proprietary files, etc.)
https://www.kai-waehner.de/blog/2020/08/07/apache-kafka-handling-large-messages-and-files-for-image-video-audio-processing/
https://www.upsolver.com/blog/apache-kafka-use-cases-when-to-use-not
https://kafka.apache.org/intro
https://docs.confluent.io/kafka-clients/librdkafka/current/overview.html
https://docs.confluent.io/platform/current/clients/librdkafka/html/md_INTRODUCTION.html
Why gettimeofday() is bad at timing things
I was reading a changelog from a library I love (unrelated but here is the link: https://liballeg.org/ ) and I stumble upon that change:
“Use clock_gettime with CLOCK_MONOTONIC instead of gettimeofday (check-switch-26)”
Then I did a bit of research on the topic, and found a good explanation here:
https://blog.habets.se/2010/09/gettimeofday-should-never-be-used-to-measure-time.html
TLDR: if you use gettimeofday to time things then your program may be affected by time shift, because gettimeofday is not monotonic. if you do not care about the date and only about elapsed time, use clock_gettime.
HackRF Portapack Mayhem Firmware, Search/Freqman evolution branch
On top of the existing ‘search and stay X seconds after a match’ the SearchApp have been updated with a ‘search and stay’ mode: stay while matching, leave after X seconds of inactivity, reset counters on activity during the wait. mode,
To use it just specify a negative value in the ‘wait’ field using the rotary encoder.
As some have some problems getting the bin back from discord, here you are:
The documentation is on the wiki, here:
Learn How To Code
A lot of us already know the excellent https://openclassrooms.com/fr/
There is also, in a more game dev oriented way, https://www.codingame.com/start or https://codecombat.com/
Learning is fun, but it it even funnier if you’re learning while gaming 😉